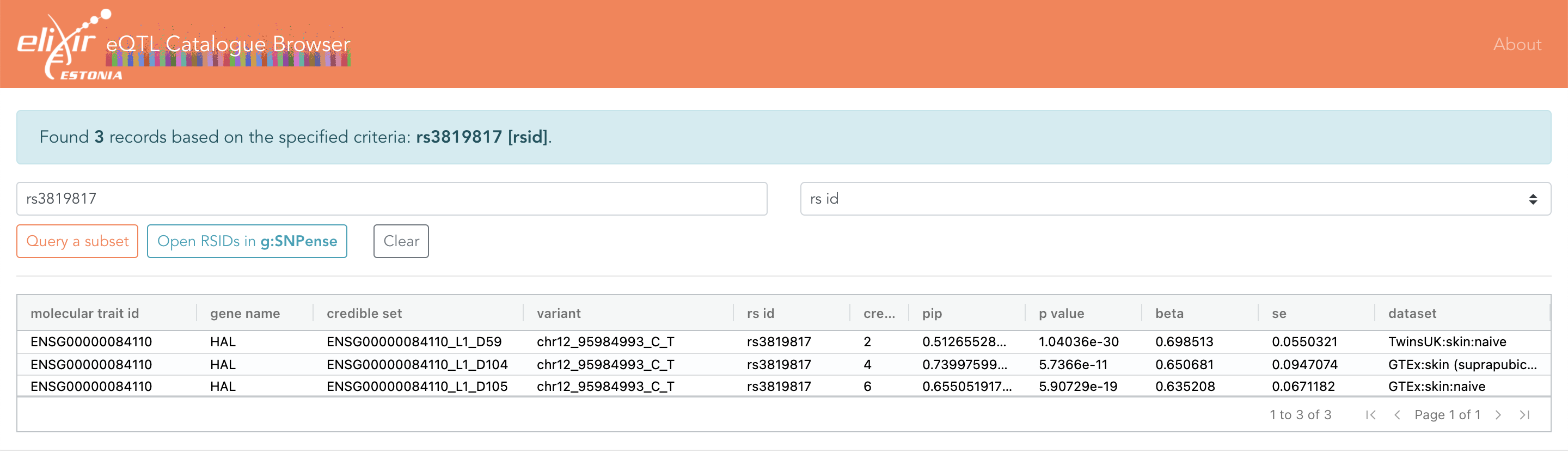
Eqtl Catalogue
Eqtl Catalogue - The four primary workflows are: Eqtl catalogue currently contains >100 datasets from 20 different studies. The eqtl catalogue provides uniform processing of 21 eqtl studies, allowing the identification of highly reproducible eqtls affecting whole gene and transcript expression levels. It enables the integration, fine mapping and. Rapid querying, colocalization, and plotting of summary stats from the eqtl catalogue. The eqtl catalogue is an open database of uniformly processed human molecular quantitative trait loci (qtls). You can access the results via ftp, api or open targets genetics portal. We are continuously updating the resource to further. Currently hosting eqtl and sqtl data from 16 different studies from the ebi eqtl catalogue 29 rows the eqtl catalogue is a database of expression quantitative trait loci (eqtl). All data analysis for the eqtl catalogue is performed using reproducible and containerised nextflow workflows. Eqtl catalogue currently contains >100 datasets from 20 different studies. The eqtl catalogue provides uniform processing of 21 eqtl studies, allowing the identification of highly reproducible eqtls affecting whole gene and transcript expression levels. Currently hosting eqtl and sqtl data from 16 different studies from the ebi eqtl catalogue 29 rows the eqtl catalogue is a database of expression quantitative trait loci (eqtl). The eqtl catalogue is an open database of uniformly processed human molecular quantitative trait loci (qtls). We are continuously updating the resource to further. Rapid querying, colocalization, and plotting of summary stats from the eqtl catalogue. The eqtl catalogue is a new project aimed at accelerating the functional interpretation of human gwas studies and the annotation of common variants through a comprehensive collection of. The four primary workflows are: The eqtl catalogue is a compendium of uniformly processed eqtls from 21 studies, covering various cell types and tissues. The eqtl catalogue provides uniform processing of 21 eqtl studies, allowing the identification of highly reproducible eqtls affecting whole gene and transcript expression levels. It enables the integration, fine mapping and. You can access the results via ftp, api or open. The eqtl catalogue is a new project aimed at accelerating the functional interpretation of human gwas studies and the annotation of common variants through a comprehensive collection of. Currently hosting eqtl and sqtl data from 16 different studies from the ebi eqtl catalogue You can access the results via ftp, api or open targets genetics portal. 29 rows the eqtl. The eqtl catalogue is an open database of uniformly processed human molecular quantitative trait loci (qtls). The four primary workflows are: All data analysis for the eqtl catalogue is performed using reproducible and containerised nextflow workflows. The eqtl catalogue is a compendium of uniformly processed eqtls from 21 studies, covering various cell types and tissues. The eqtl catalogue provides uniform. The eqtl catalogue is a compendium of uniformly processed eqtls from 21 studies, covering various cell types and tissues. The eqtl catalogue is a new project aimed at accelerating the functional interpretation of human gwas studies and the annotation of common variants through a comprehensive collection of. We are continuously updating the resource to further. You can access the results. It enables the integration, fine mapping and. Eqtl catalogue currently contains >100 datasets from 20 different studies. The eqtl catalogue is a compendium of uniformly processed eqtls from 21 studies, covering various cell types and tissues. Rapid querying, colocalization, and plotting of summary stats from the eqtl catalogue. The eqtl catalogue is an open database of uniformly processed human molecular. Eqtl catalogue currently contains >100 datasets from 20 different studies. The eqtl catalogue provides uniform processing of 21 eqtl studies, allowing the identification of highly reproducible eqtls affecting whole gene and transcript expression levels. The eqtl catalogue is an open database of uniformly processed human molecular quantitative trait loci (qtls). The eqtl catalogue is a compendium of uniformly processed eqtls. The eqtl catalogue is a new project aimed at accelerating the functional interpretation of human gwas studies and the annotation of common variants through a comprehensive collection of. You can access the results via ftp, api or open targets genetics portal. It enables the integration, fine mapping and. Eqtl catalogue currently contains >100 datasets from 20 different studies. Rapid querying,. The eqtl catalogue provides uniform processing of 21 eqtl studies, allowing the identification of highly reproducible eqtls affecting whole gene and transcript expression levels. The eqtl catalogue is an open database of uniformly processed human molecular quantitative trait loci (qtls). 29 rows the eqtl catalogue is a database of expression quantitative trait loci (eqtl). Rapid querying, colocalization, and plotting of. We are continuously updating the resource to further. The eqtl catalogue provides uniform processing of 21 eqtl studies, allowing the identification of highly reproducible eqtls affecting whole gene and transcript expression levels. It enables the integration, fine mapping and. Currently hosting eqtl and sqtl data from 16 different studies from the ebi eqtl catalogue 29 rows the eqtl catalogue is. The eqtl catalogue is an open database of uniformly processed human molecular quantitative trait loci (qtls). Eqtl catalogue currently contains >100 datasets from 20 different studies. The eqtl catalogue provides uniform processing of 21 eqtl studies, allowing the identification of highly reproducible eqtls affecting whole gene and transcript expression levels. It enables the integration, fine mapping and. The eqtl catalogue. It enables the integration, fine mapping and. The eqtl catalogue is an open database of uniformly processed human molecular quantitative trait loci (qtls). Currently hosting eqtl and sqtl data from 16 different studies from the ebi eqtl catalogue The eqtl catalogue is a compendium of uniformly processed eqtls from 21 studies, covering various cell types and tissues. The eqtl catalogue provides uniform processing of 21 eqtl studies, allowing the identification of highly reproducible eqtls affecting whole gene and transcript expression levels. The eqtl catalogue is a new project aimed at accelerating the functional interpretation of human gwas studies and the annotation of common variants through a comprehensive collection of. All data analysis for the eqtl catalogue is performed using reproducible and containerised nextflow workflows. 29 rows the eqtl catalogue is a database of expression quantitative trait loci (eqtl). The four primary workflows are: Rapid querying, colocalization, and plotting of summary stats from the eqtl catalogue.Overview of the datasets included in the eQTL Catalogue. (A
GitHub eQTLCatalogue/eQTLCatalogueresources
Building the eQTL Catalogue a Q&A with Kaur Alasoo
Overview of studies and samples included in the eQTL Catalogue a
(PDF) eQTL Catalogue a compendium of uniformly processed human gene
Overview of the eQTL Catalogue database a, A highlevel representation
(PDF) eQTL Catalogue 2023 New datasets, X chromosome QTLs, and
Overview of the eQTL Catalogue database a, A highlevel representation
Building the eQTL Catalogue a Q&A with Kaur Alasoo
eQTL Catalogue a new resource for gene expression and splicing QTLs
Eqtl Catalogue Currently Contains >100 Datasets From 20 Different Studies.
You Can Access The Results Via Ftp, Api Or Open Targets Genetics Portal.
We Are Continuously Updating The Resource To Further.
Related Post: